SPADA is a CRISPR-based genome editing system that uses engineered viruses to efficiently and specifically disrupt multiple genes in targeted brain cells, enabling precise modeling of complex genetic brain disorders in animals.
Background
The field of neurogenetics relies heavily on genome editing to understand the mechanisms of complex psychiatric illnesses, such as bipolar disorder. Many brain disorders are polygenic, driven by the combined effects of multiple genetic variants rather than a single mutation. To accurately model these conditions and dissect the complex genetic architecture identified by Genome-Wide Association Studies, researchers need tools capable of manipulating specific neural circuits in vivo. There is a critical need for highly efficient, cell-type-specific genome editing methods that can simultaneously disrupt multiple genetic loci to reflect the multigenic nature of these diseases.
Current genome editing approaches face significant limitations when attempting to model multigenic conditions. Traditional single-guide CRISPR methods rely on stochastic insertions or deletions to disrupt genes. This often results in unpredictable in-frame mutations and residual protein expression, failing to achieve a reliable loss-of-function. Furthermore, delivering multiple editing components to post-mitotic neurons is severely restricted by the strict packaging limits of standard viral vectors, making simultaneous multi-locus editing highly challenging. Finally, optimizing guide RNAs directly in animal models without predictive in vitro screening is a slow, expensive process requiring many research animals.
Technology Description
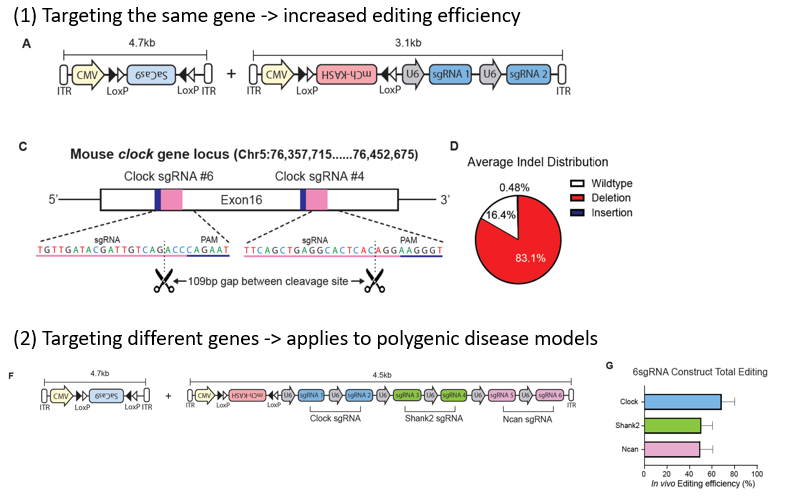
The SPADA platform is a cell-type-specific genome editing solution designed for post-mitotic neurons. It utilizes a two-vector adeno-associated virus (AAV) system: one vector delivers a Cre-dependent Staphylococcus aureus Cas9, while the second carries a nuclear reporter and up to six single guide RNAs (sgRNAs) (Figure 1). A core feature is its dual-guide excision strategy, targeting two sites approximately 100 base pairs apart within a single gene to produce defined genomic deletions. The system supports extensive multiplexing to simultaneously disrupt up to three independent genetic loci and incorporates a predictive in vitro-to-in vivo screening pipeline to optimize sgRNA selection.
This technology is differentiated by its ability to overcome the limitations of traditional single-guide CRISPR approaches, which often rely on unpredictable mutations that leave residual protein expression. By employing the dual-guide excision method, the platform guarantees precise genomic excisions and frameshift mutation rates exceeding 90%, ensuring a robust and predictable loss-of-function. Additionally, its unique multiplexing capability allows researchers to accurately model complex polygenic conditions, such as psychiatric illnesses, by simultaneously targeting multiple genes within specific neural circuits in vivo. Finally, the predictive screening pipeline significantly reduces the time, cost, and animal use required for research.
Benefits
- Achieves highly efficient and precise gene knockouts with over 90% frameshift mutation rates via a dual-guide excision strategy, ensuring robust and predictable loss-of-function.
- Enables the simultaneous disruption of up to three independent genetic loci by multiplexing up to six sgRNAs in a single construct, making it ideal for modeling complex polygenic diseases.
- Provides precise, cell-type-specific genome editing through Cre-dependent expression, allowing targeted modifications in specific cell populations such as post-mitotic neurons.
- Overcomes restrictive viral packaging limits for efficient in vivo delivery by utilizing a compact SaCas9 enzyme within a two-vector AAV system.
- Reduces research time, costs, and animal usage through a predictive in vitro-to-in vivo pipeline that allows for rapid screening and optimization of guide RNAs.
- Accelerates the development of custom disease models and knockout animal lines, facilitating the discovery and screening of novel therapeutic targets for complex disorders.
Commercial Applications
- Custom polygenic disease modeling
- Multiplexed therapeutic target screening
- Knockout animal line production
- Neurological gene therapy development
- Psychiatric drug discovery modeling
Additional Information
This two-vector AAV platform enables cell-type-specific, multiplexed genome editing in neurons. It utilizes a Cre-dependent SaCas9 and a vector delivering up to six sgRNAs. A dual-guide excision strategy targeting sites 100 base pairs apart achieves frameshift mutation rates exceeding 90%. The system supports simultaneous disruption of three independent genetic loci, facilitating high-efficiency modeling of complex polygenic conditions through precise, scalable in vivo genomic deletions.

